DBBS supports various optional, prestigious ‘value added,’ cross-programmatic
curricular and co-curricular experiences that support interdisciplinary trainee
research and interests. Some of these enrichment opportunities come with
fellowships that cover or augment stipends.
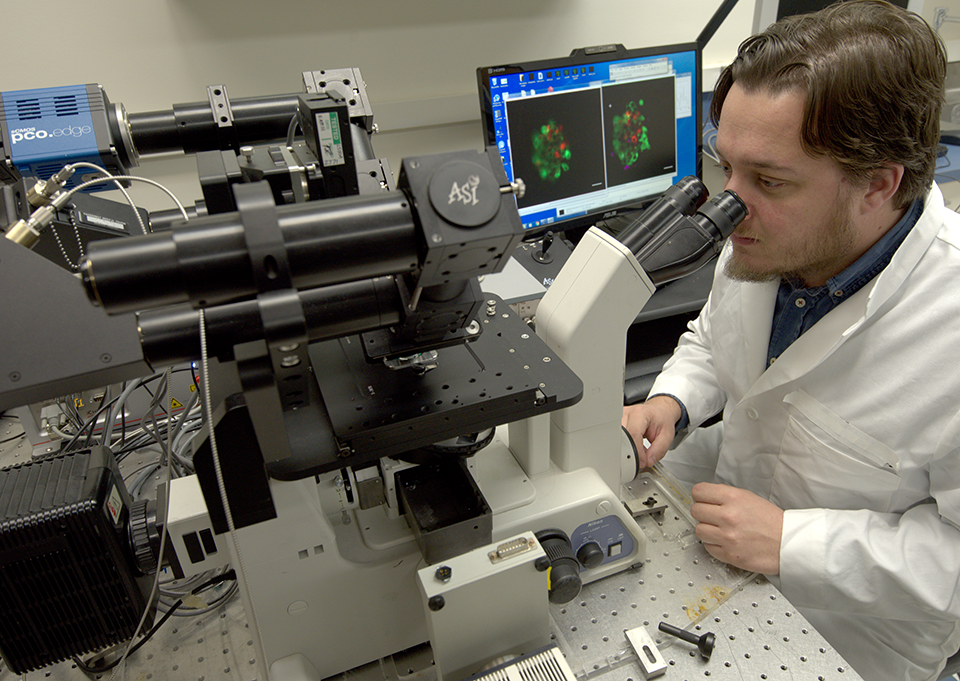

Cancer Biology Pathway
Distinct from the degree-granting Cancer Biology PhD program, this NIH-funded pathway is for second- and third-year students from any PhD training program (MSTP students are not eligible). Students should be working on a cancer biology research problem and interested in enriching their curriculum in this area.

Cells to Society Pathway
The Cells to Society Pathway brings rigorous training in biological sciences, biostatics/statistical genomics and epidemiology together to foster leadership in biological and quantitative population sciences, applied particularly to problems in cancer biology. The pathway will transform graduate training and expedite the move from bench to bedside by emphasizing a population perspective on cancer prevention and treatment. Students enter this pathway in year one of their graduate training.
Contact Graham Colditz or Susan Dutcher for more information.

Chemical Biology
MSTP and DBBS students are invited to consider enrichment in Chemical Biology, a modern field with roots in pharmacology, bioorganic chemistry, biochemistry, synthetic biology, proteomics, and molecular imaging. Medicinal chemistry and drug discovery are also important components of Chemical Biology training. Students interested in pursuing this area of research can train with Chem Bio-adjacent faculty, many of whom are listed on the fact sheet link below. Students who complete either Medicinal Chemistry I (CHEM5571) or Introduction to Biomolecules (Chem 5811) and Chemical Biology (CHEM5821) will earn an Emphasis in Chemical Biology, including a certificate of completion and an electronic micro credential. Other courses with related subject
matter are also available.
Learn more about the Chemical Biology Enrichment Pathway or contact Jim Janetka.

Cognitive, Computational, and Systems Neurosciences (CCSN) Pathway
One of our longest standing value-added pathways, CCSN draws students from programs in Biomedical Engineering, Psychological & Brain Sciences, and DBBS Neuroscience together for courses and workshops and fosters thesis projects at the intersection of these disciplines. Any student can take a la carte curricular offerings from CCSN, but the formal pathway starts in years one to two of the home program’s training. A limited number of training grant positions are available for students engaged in the full pathway.
Learn more about the CCSN Pathway.

Human Genetics
Students from Genetics-adjacent PhD programs such as Computational and Systems Biology, Molecular Genetics and Genomics, Bioinformatics and Data Sciences,
Neurosciences, and others may want to enhance their understanding of modern methods and approaches in human genetics. Students wishing to complete an
Emphasis in Human Genetics commit to taking two elective courses, Human Genetic Analysis (Bio5483) and Computational Statistical Genetics (MSB621). Completion is
awarded with a certificate of completion and an electronic micro credential. There is no formal admission process for the emphasis.
Interested students should contact Dr. Nancy Saccone for more information.
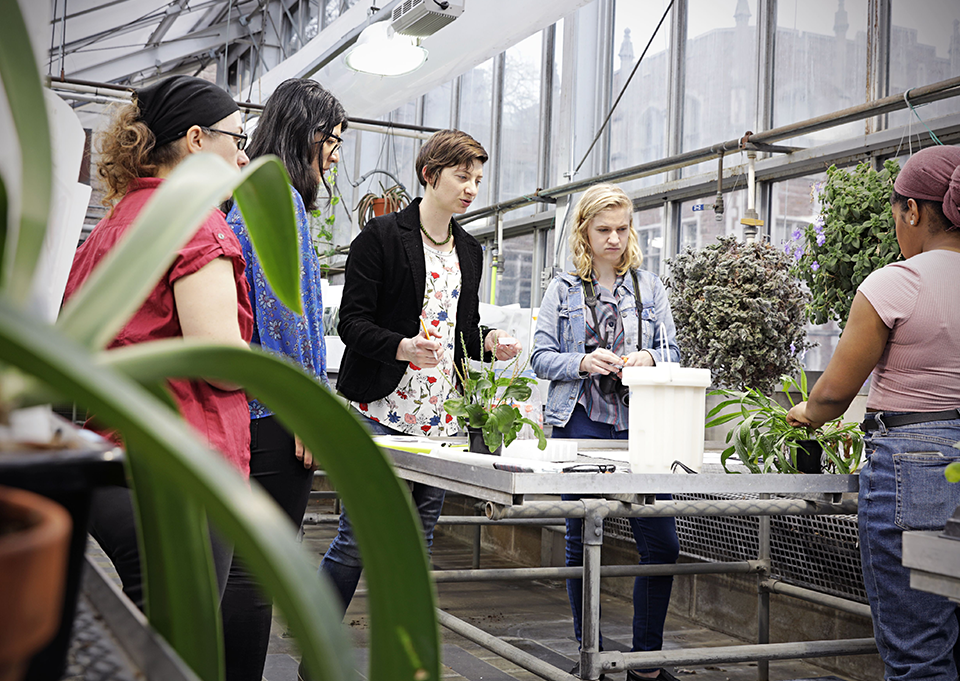

William H. Danforth Plant Sciences Pathway
The William H. Danforth Plant Sciences Fellowship seeks to foster a culture of intellectual entrepreneurship focused on research, leadership and innovation in the general area of plant science. The fellowship is open to first-year Washington University PhD students in any DBBS-affiliated laboratory with a thesis project in the general area of plant biology. Selected students will receive a 4-year fellowship to cover yearly stipend plus $2,000 additional per year for project supplies, meeting attendance and research-related travel. Danforth Fellows will complete their program courses, take the Fellows Gateway Course (advanced elective), participate in additional career development activities, and finish a PhD thesis.
Interested students should contact Joe Jez or or Ram Dixit for more information.
Precision Medicine Pathway
The Precision Medicine Pathway will introduce students to the use of genomic and genetic information in the diagnosis and treatment of disease. The pathway will expand students’ educational experience into the clinical realm; facilitate making clinical connections related to thesis research, where relevant; and provide career path information, including potential areas for postdoctoral research and non-academic routes. The pathway is available to graduate students in the second and third year who have knowledge in genetics, genomics and bioinformatics. In most cases, students come from Computational Systems Biology, Bioinformatics and Data Sciences, or Molecular Genetics & Genomics DBBS programs. This knowledge base will allow lectures and discussions to be at an advanced level in genetics and genomics, highlighting clinical aspects.
Learn more about the Precision Medicine Pathway or contact Dr. Tim Schedl.
Imaging Sciences Pathway
The Imaging Sciences Pathway is open to students in graduate programs in DBBS, Chemistry, Physics and the School of Engineering and Applied Sciences. With over 30 mentors from 11 departments on the Danforth and medical campuses, the pathway offers diverse, multidisciplinary opportunities. Career trajectories of graduates may include imaging technology development, the chemistry and use of novel contrast agents, the visualization and manipulation of macromolecular complexes and organelles in cells and in animals, and the application of these technologies to the visualization of human disease states. Students’ education will be enhanced by interdisciplinary coursework and research experiences in Imaging Sciences, starting in their second year. A limited number of stipend fellowships are available.
Learn more about the Imaging Sciences Pathway or contact Joe Culver for more information.
The Infectious Disease Gateway
The Infectious Disease Gateway is a single value-added course that provides an opportunity for students, postdoctoral fellows and infectious disease fellows to explore issues at the interface between patient care, public health and basic research in the area of microbial pathogenesis. It provides a glimpse of alternative career pathways beyond that of a PI in an academic environment. The ID Gateway course is open to PhD & MSTP students and postdoctoral fellows in any department or program. Graduate and MSTP students are expected to have successfully completed either the graduate or medical microbiology course, although this requirement may be waived at the discretion of the course directors. Qualified students apply in the fall for notification in time for registration for spring semester.
Contact Michelle Potter for more information.
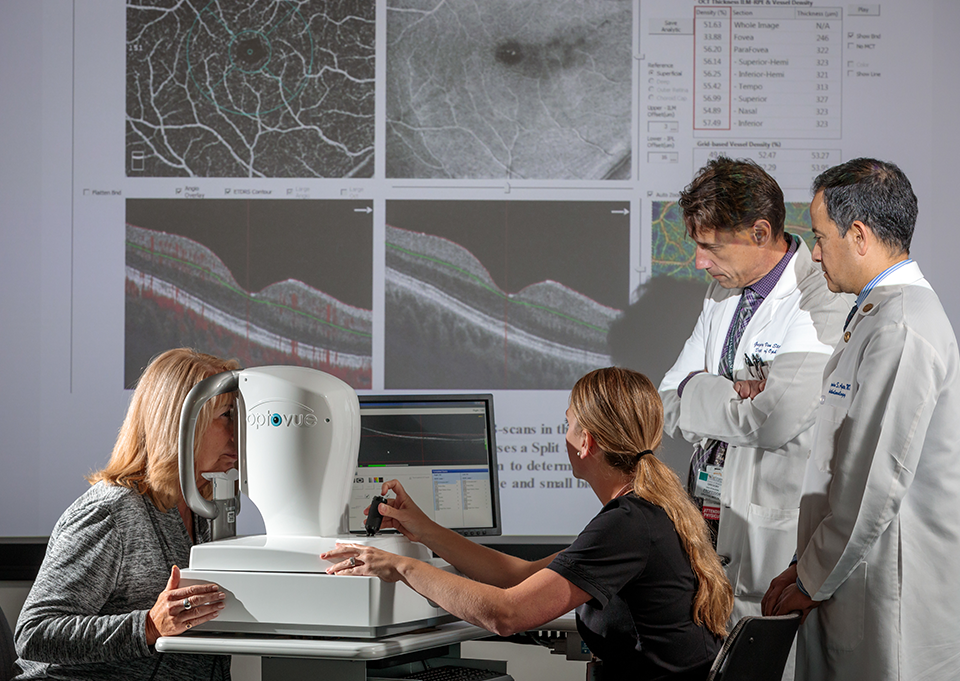

The Interdisciplinary Training in Vision Science (ITVS) Pathway
The ITVS Pathway is a 2-year program that overcomes boundaries between disciplines to develop a unique set of basic science, translational, and professional skills that will position students to produce transformative breakthroughs in vision science. Completion of the pathway entails 4 courses that introduce the biology and pathology of the visual system, translating basic science to clinical applications, advanced techniques used to study the visual system, and project building/support for grant writing. Participation in the pathway qualifies students for up to 2 years of T32 funding support and to compete for the annual Winston Fellow Award that recognizes outstanding contributions to vision science.
Learn more about the ITVS Pathway or contact Jenna Krizan for more information.
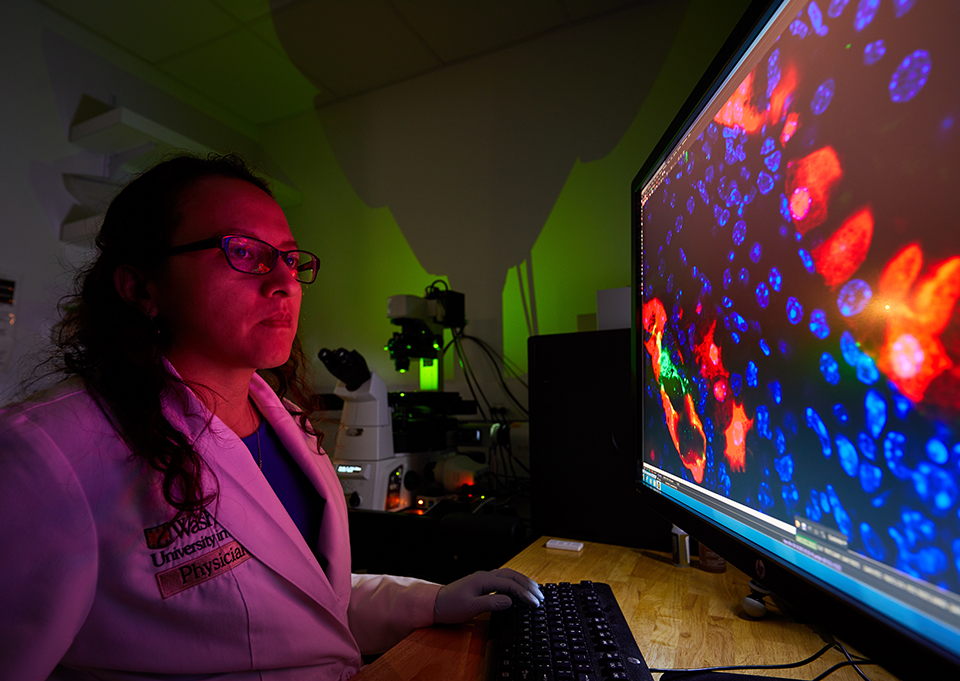

Lucille P. Markey Special Emphasis Pathway in Human Pathobiology
Although the primary mission of DBBS is training in foundational biosciences, the Markey Pathway helps bridge the gap between foundational and clinical research. The pathway trains DBBS students in various aspects of human disease not generally covered in graduate courses. The Markey Pathway is a 2-year course of study including Pathobiology of Human Disease States Course, Individual Clinical Mentorship, and an Annual Retreat. Each year 8-10 students with at least two years left in their graduate careers are selected as Markey Students. A one-time stipend supplement of $4,000 will be provided during the first year of the Pathway to accepted students’ PI.
Learn more about the Markey Pathway, or contact markeypathway@kids.wustl.edu for more information.